This website provides informations about LibGEP, a C++ library dedicated to Gene Expression Programming.
This page proposes an overview of the following elements:
More precisely, GEP encode chromosomes as symbolic string of fixed length composed of one or more genes. This string contains two kinds of symbols: functions (from a set F) and terminals (belonging to a set T).
For instance, let F = {+,-,*,√} and T = {a,b,c}. Figure 1 provides the gene and the expression tree associated to the algebraic expression (a+b)*√(b - c). Genes in GEP are composed of a head (containing both functions and terminals) together with a tail containing only terminals. Let h (resp. t) be the size of the head (resp. the tail). Then GEP encode each gene with a string of length h+t (t = h(a - 1) + 1) where a is the maximum number of arguments taken by functions in F (in our example, a = 2) and t = h + 1 such that each gene is encoded with a string of length 2h+1. For instance, let's take h=7. Then the complete gene considered previously in Figure 1 could be written in GEP :
In this case, the expression tree finishes at position 7 whereas the gene ends at position
14.
If a mutation occurs at position 4 that change 'b' into '+', then the following gene
is obtained:
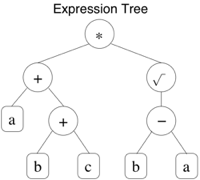
The corresponding expression tree is given in Figure 2 (and lead to the function (a+b+c)*√(b-a)). It now finishes at position 9.
The way cross-over operates is similar.
So despite its fixed length, each gene has the potential to code for expression trees
of different sizes and shapes.
Later on, Sebastien joined the project as he was looking for a framework to analyse cheating impact on evolutionary processes. This came with a full rewrite in C++ and a better packaging through autotools.
So to sum up, LibGEP is a C++ library dedicated to Gene Expression Programming (GEP), highly configurable through the massive use of templates.
LibGEP is still in beta (latest release: 0.4).
This is an open-source project yet if you use it, it would be nice to:
- Evolutionary Algorithms
- What is Gene Expression Programming?
- Development Status and History
- Licence
- Authors
Evolutionary Algorithms
Evolutionary Algorithms (EAs) is a class of solving technique based on the Darwinian theory of evolution which involve the search of a population of solutions and not only one. A possible and acceptable solution i.e. a member of the population is called an individual. Each iteration of an EA involves a competitive selection that weeds out poor solutions through the evaluation of a fitness value that indicate the quality of the individual as a solution to the problem. The evolutionnary process involves at each generation a set of genetic operators that are randomly applied on the individuals, typically recombination (or cross-over) and mutation.
(top)
What is Gene Expression Programming?
John Koza in [Koza_GP] proposed to use Genetic Algorithm (GA) in so called genetic programming (GP). Gene expression programming (GEP) was proposed by Candida Ferreira in [Ferreira_GEP01] as an extension of GP. As an EA, GEP use populations of individuals, select the individuals according to fitness, and introduce genetic variation using one or more genetic operators. The fundamental difference between these three classes of EAs resides in the nature of the individuals:- in GAs the individuals are symbolic strings of fixed length (chromosomes);
- in GP the individuals are nonlinear entities of different sizes and shapes called
parse trees (or expression tree) that represent a program/function (see Figure 1);
Figure 1: GP genes and the associated expression tree. 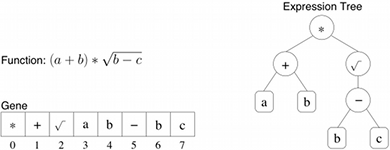
- in GEP the individuals are also nonlinear entities of different sizes and shapes (expression trees), but these complex entities are encoded as simple strings of fixed length (chromosomes).
More precisely, GEP encode chromosomes as symbolic string of fixed length composed of one or more genes. This string contains two kinds of symbols: functions (from a set F) and terminals (belonging to a set T).
For instance, let F = {+,-,*,√} and T = {a,b,c}. Figure 1 provides the gene and the expression tree associated to the algebraic expression (a+b)*√(b - c). Genes in GEP are composed of a head (containing both functions and terminals) together with a tail containing only terminals. Let h (resp. t) be the size of the head (resp. the tail). Then GEP encode each gene with a string of length h+t (t = h(a - 1) + 1) where a is the maximum number of arguments taken by functions in F (in our example, a = 2) and t = h + 1 such that each gene is encoded with a string of length 2h+1. For instance, let's take h=7. Then the complete gene considered previously in Figure 1 could be written in GEP :
| head | tail |
| * + √ a b - b | c b a a a b c b |
| head | tail |
| * + √ a + - b | c b a a a b c b |
 |
(top)
Development Status and History
It appeared difficult to find an open-source library that makes it possible to use GEP for a variety of programs (actually, a commercial product is available here). Marek wanted to apply this approach for some function finding and therefore decided to implement its own version (initially in Java) on the basis of [Ferreira_GEP01] and [Ferreira_GEP06].Later on, Sebastien joined the project as he was looking for a framework to analyse cheating impact on evolutionary processes. This came with a full rewrite in C++ and a better packaging through autotools.
So to sum up, LibGEP is a C++ library dedicated to Gene Expression Programming (GEP), highly configurable through the massive use of templates.
LibGEP is still in beta (latest release: 0.4).
(top)
Licence
LibGEP is released under GNU GPL Licence v3 (free software).
(top)
Authors
LibGEP is written by Marek Ostaszewski and Sebastien Varrette.This is an open-source project yet if you use it, it would be nice to:
- inform us (with a short description of the way you use LibGEP) and
- give us some feedback about the library (see Submit Bugs for instance)


